By Amy Scott, PhD Candidate, Department of Anthropology, Boston University
“One, two, three, four, five, six, seven, eight, one, two, three, four, five, six, seven, eight, satu, dua, tiga, empat, lima, enam, tujuh, delapan…” I count to myself as I fill tubes with chemicals, occasionally counting in Indonesian to keep my mind active. Since October, I have been conducting genetics research at the Eijkman Institute for Molecular Biology in Jakarta to determine the paternity of orangutans born in Gunung Palung National Park. We want to know if flanged or unflanged males sire more offspring. We also want to know if one flanged male is able to monopolize paternity. Because orangutans are primarily solitary and females mate with multiple males, genetic analysis is the only reliable method for determining paternity. This is part of my dissertation research at Boston University, examining reproductive strategies of orangutans. My excitement to learn the results motivates me throughout the monotonous task of carefully counting and filling multiple tubes with different chemicals all day long!
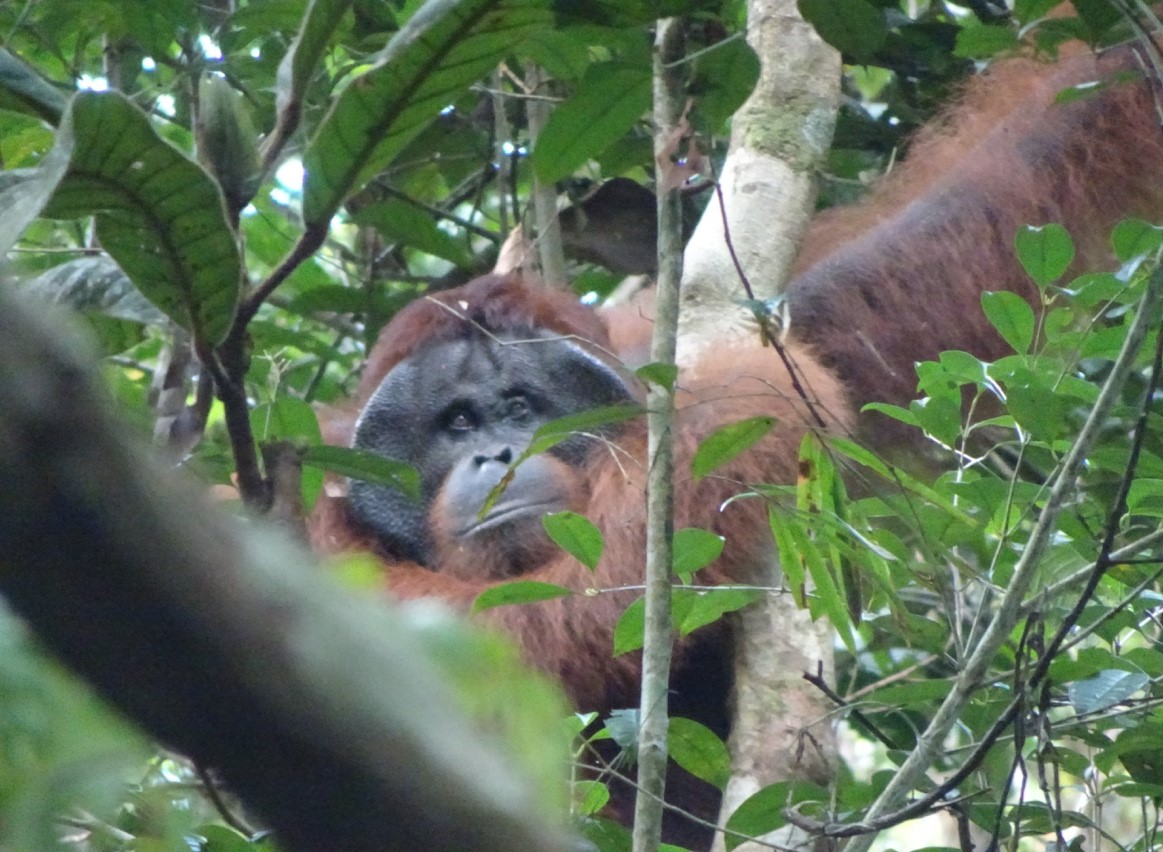
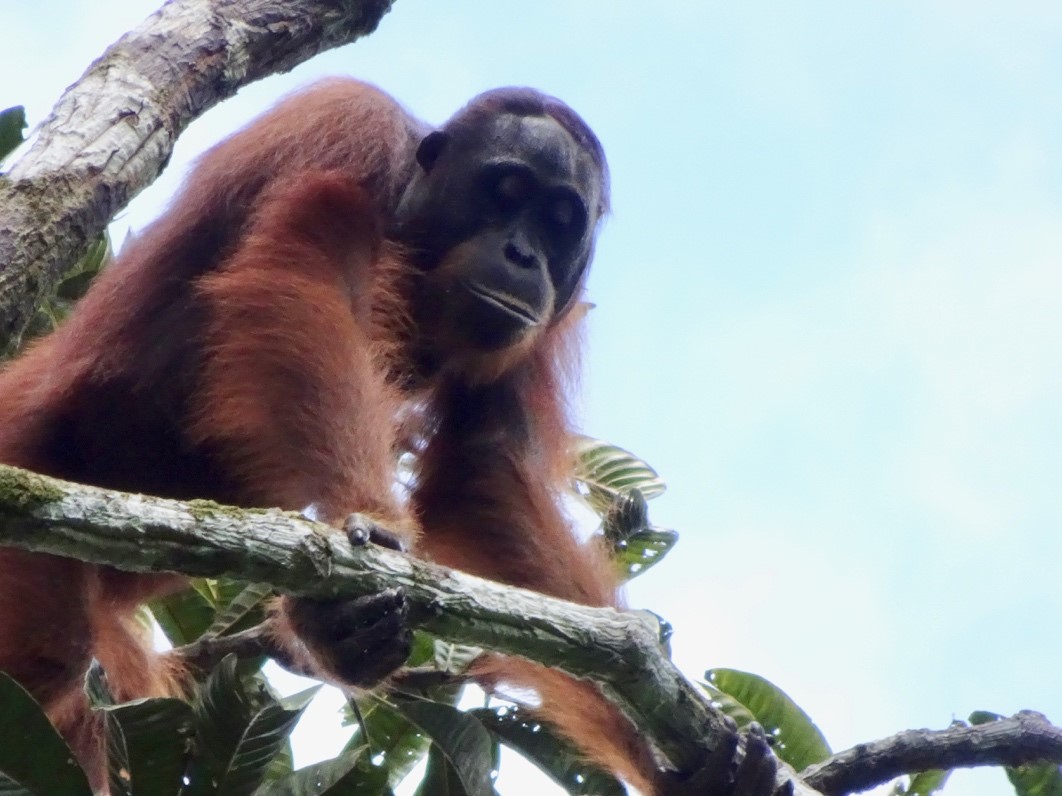


For this project, I am processing orangutan fecal samples that have been collected by dozens of field assistants, students, volunteers, and managers at Cabang Panti since 2008. Fecal samples are the best non-invasive source of DNA that we can collect from wild orangutans. It is important that our research is purely observational and does not harm any orangutans, so collecting a blood, hair, saliva or tissue sample from our wild orangutan study population is not an option. However, the only drawback to fecal samples is that they only contain small amounts of DNA compared to invasive samples, like blood. The fecal samples are stored in special chemicals or dried to protect the DNA from degradation, and then transported to freezers in Ketapang, and eventually brought to freezers at the Eijkman Institute in Jakarta.
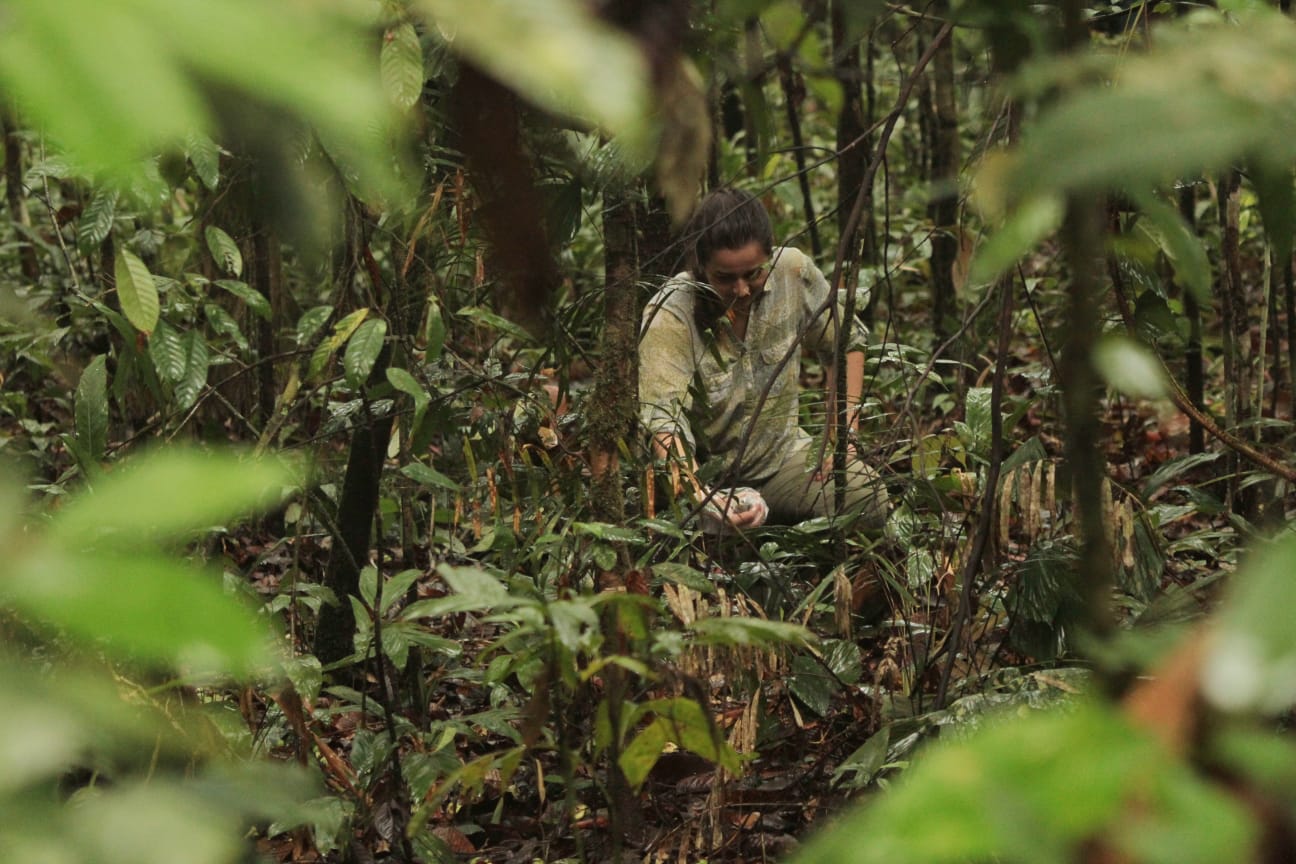

In order to determine paternity, a sample is needed from the offspring, the mother, and all possible male sires. I will genotype each of these animals. Genotyping is like fingerprinting, but with DNA. I am looking at 12 different locations in the orangutan DNA. Every individual has two markers (alleles) at each location, and there are 4-10 possible markers (alleles) at each of those locations. This many different combinations of markers means that each individual will have a unique combination of markers, i.e. each individual has a unique genetic fingerprint. Just like all other sexually reproducing animals, half of the offspring’s DNA comes from the mother and half from the father.
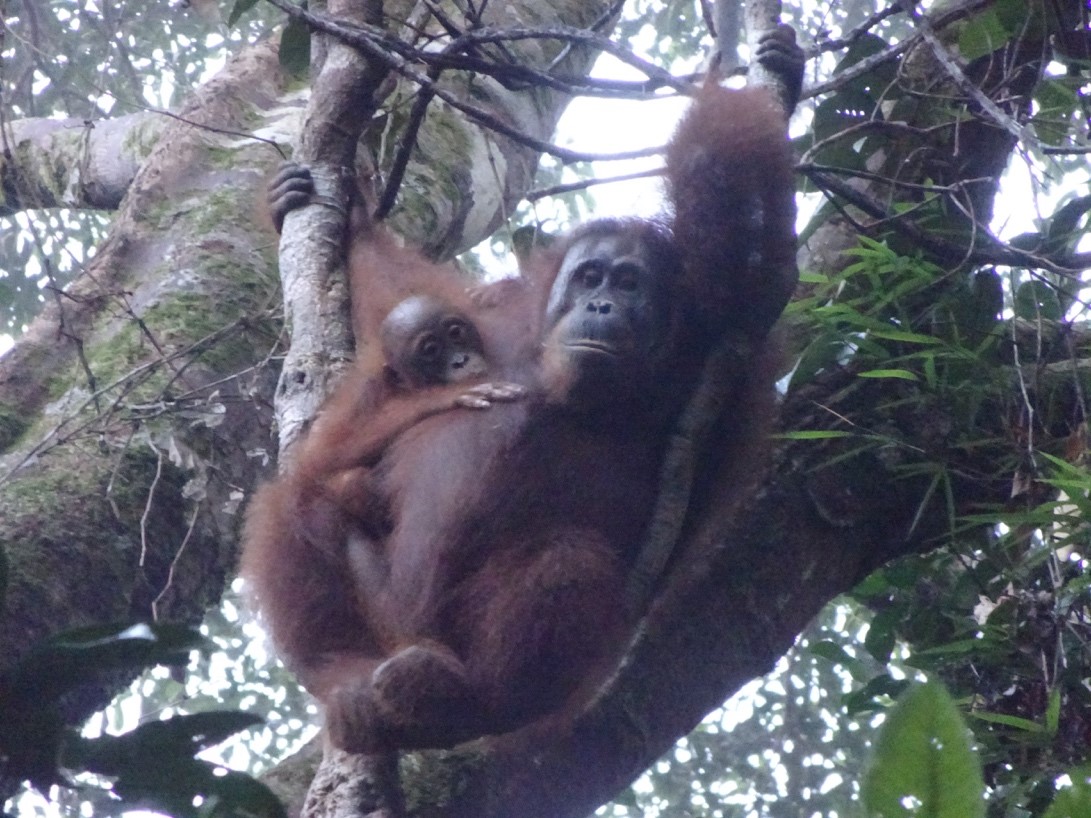

Since we know from behavioral observations who the mother of the offspring is, we can identify which one of the pair of markers at each location in the DNA comes from the mother. By process of elimination, the other marker must come from the father. This will tell us which markers the father must have at each of the 12 locations. Next, we can compare this genetic signature of the offspring’s father with the genotypes of all the potential sires, and hopefully find the match. We will be using a computer program to make these comparisons.
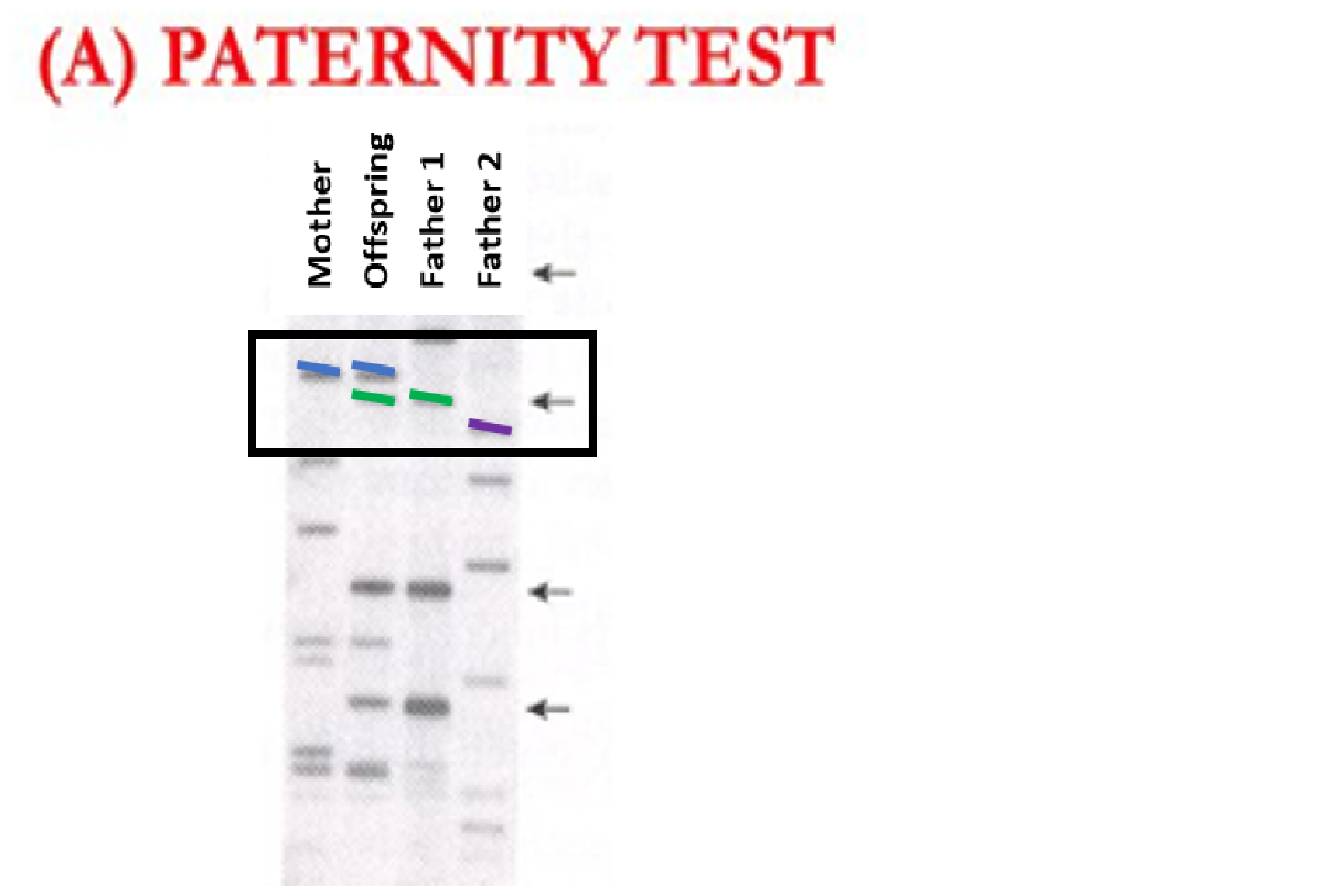
The above example shows genotypes from 4 individuals. Individuals always have two markers at a given location in their DNA, so if they have two copies of the same marker (homozygous) it will show up as a single marker. If an individual has two different markers (heterozygous) it will show two different colored markers. At the location in the DNA that is signified with a box around it, we see that the mother has two copies of the blue marker (homozygous) and the offspring has a blue and a green marker (heterozygous). Because the mother will pass a blue marker on to all of her offspring, the offspring’s blue marker must come from the mother. Therefore, the offspring’s green marker must come from the father. Because only Father 1 has green markers at this location and Father 2 does not have green markers, Father 1 is a candidate father whereas Father 2 is not (color added for clarity, in reality these markers are differentiated based on length.)
Genotyping the orangutans requires four steps in the lab: extraction, quantification, amplification, and fragment analysis. First, we extract the DNA using a special kit with chemicals, beads, and heat to break up cells and pull out the DNA from inside the cells. Next, we quantify the amount of orangutan DNA that is in the sample. We need to know how much DNA is in each sample in order to know how many replicates are needed in the following steps, ensuring that the resulting genotype is correct. Due to the low amount of DNA in fecal samples, the next steps are prone to errors, but if you replicate them (up to 5 times), you can ensure 99% accuracy of genotyping. The third step is PCR (Polymerase chain reaction) which amplifies the DNA. Machines called thermocyclers make many copies of the specific locations in the DNA that we are interested in. We repeat this step two times to increase accuracy. Then finally, we conduct fragment analysis to see what markers each orangutan has at the 12 locations. This final step requires special computer software to read how long the markers are for each location in each individual.

Each one of these steps of lab work is very intricate with many detailed steps that must be carefully followed. As with all lab work, there is trial and error to optimize the best conditions and combinations. Still, the most important thing is carrying out each step carefully. With genotyping, it is very important to keep track of each sample and not cross-contaminate, from one individual to another. I am working with a plate that has 96 wells (12 columns of 8 rows), so I have to be very careful that I am always using the correct sample and avoiding sample contamination. The repetitiveness of counting to 8 all day in an air-conditioned lab, is very different than the fieldwork portion of my dissertation research (see Code Red Volume 69, September 2018) observing orangutans from sunrise to sunset in the rainforest, never-knowing what the day might bring. Combining these two very different types of research with the long-term database will enable us to reveal the paternity of the Gunung Palung orangutans that have been born in our study area and to answer important questions about the reproductive benefits of developing flanges for male orangutans.









